DNA and RNA sequence data are being deposited in public repositories at an astonishing rate. The quantity of sequence data in GenBank has doubled approximately every 18 months since 1982. This is an extraordinarily rich source of information about microbial life.
I argue that it is impossible to use all of the information in a sequence data set in a single study, and that we can learn a great deal by combining data from disparate studies that were not initially intended to be compared.
What We’ve Done
Through bioinformatic meta-analyses, the Steen Lab has:
- Quantified uncultured microbes: Determined that high proportions of bacteria and archaea across most biomes remain uncultured
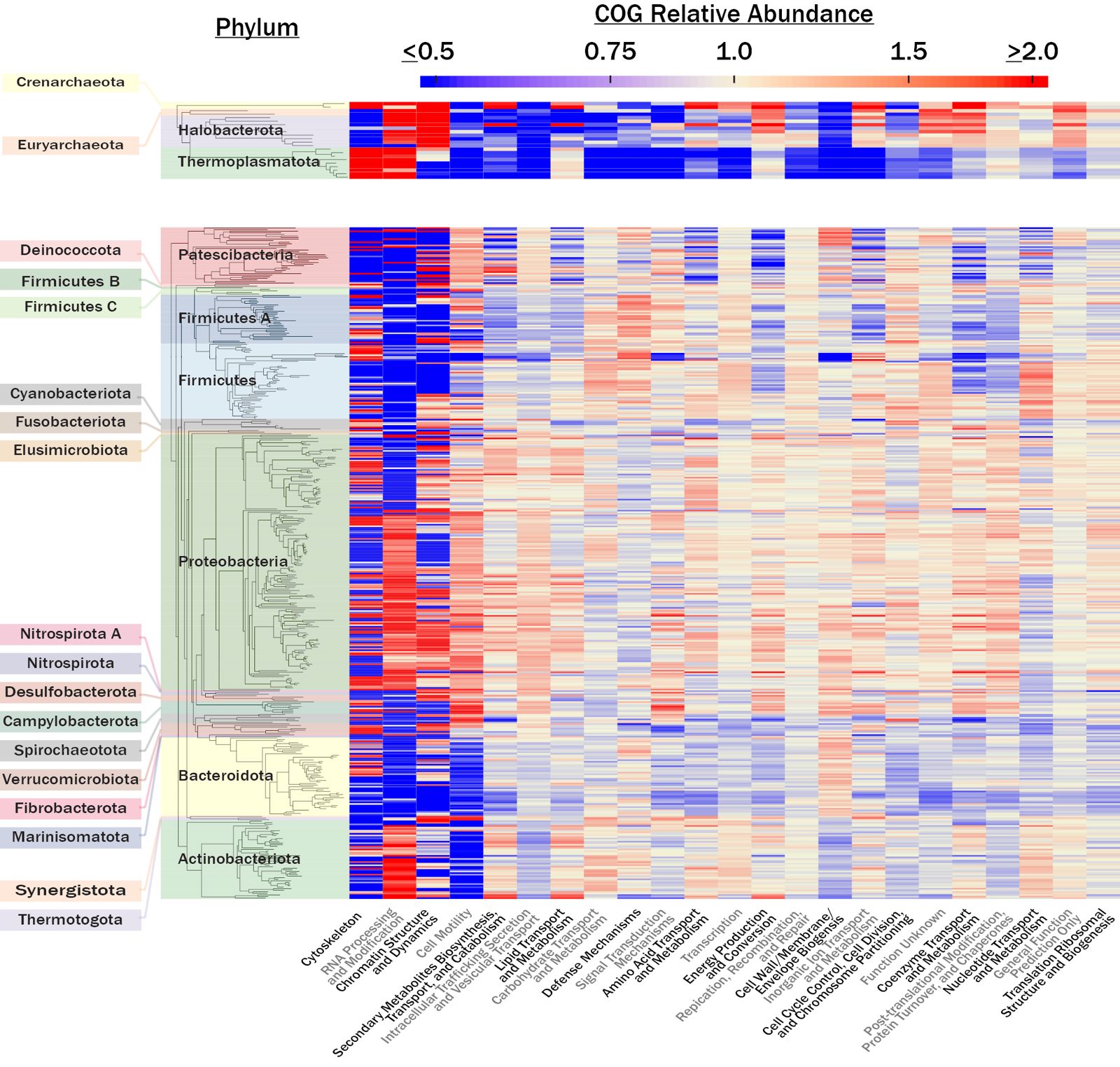
- Explored trait conservation: Studied the degree to which microbial traits are conserved as a function of taxonomic rank
- Optimized metagenomics: Investigated the relationship between DNA sequencing effort and the quantity of metagenome-assembled genomes (MAGs)
- Developed ML tools: Built a deep learning-based alignment-free sequence similarity search tool
Our approach treats public databases as an untapped scientific resource — one that can reveal fundamental truths about the structure and function of life on Earth.